更年期…自分にはいつか来るものと覚悟をしていたつもりでしたが、情緒不安定や異様に汗をかくなどこんなにも辛いものだと思っていませんでした。症状が出るとまたイライラして…というのを繰り返していてかなり参っていたときに義母に進められたのがこのプロギノバでした。1日1回のむだけで症状がかなり抑えられています。薬に頼りたくないという人もいると思うけど、辛いのを我慢するよりかは薬に頼ってもいいってことを教えてあげたいです。

左記クレジットカード、銀行振込、コンビニ決済に対応



更新日:2025/6/3
プロギノバは、更年期ならではのイライラや異常な発汗、ほてりなどの症状(更年期障害)を改善に導く医薬品です。
有効成分が崩れがちなホルモンバランスを正常に近づけることで、諸症状を緩和させます。
また、加齢によってリスクが高まる骨粗しょう症を予防する効果も発揮します。
| メーカー | バイエル(Bayer) ザイダス・カディラ(Zydus Cadila) |
|---|---|
| 有効成分 | エストラジオール吉草酸エステル |
| 効果 | 更年期障害の改善 閉経後骨粗鬆症の予防 |
| 副作用 | 頭痛や乳房痛など |
| 用法 | 毎日一定の時刻に1回分を服用 |
エストラジオール吉草酸エステルは、体内で卵胞ホルモン(エストロゲン)として働く有効成分です。更年期になると卵胞ホルモンが不足しやすく、ホルモンバランスの乱れから心身の症状が起こり、骨の新陳代謝にも影響を与えて骨粗しょう症のリスクが高まります。そんな中、エストラジオール吉草酸エステルによって卵胞ホルモンを補うことで、ホルモンバランスが回復し、諸症状が改善・予防されます。
プロギノバには、1錠あたり2mgのエストラジオール吉草酸エステルが配合されています。

| 個数 | 販売価格(1錠あたり) | 販売価格(箱) | ポイント | 購入 |
|---|---|---|---|---|
| 84錠 | 107円 | 9,060円 | 271pt | |
| 168錠 | 86円 | 14,560円 | 436pt | |
| 252錠 | 80円 | 20,160円 | 604pt |






①1万円以上で送料無料
1回の注文で10,000円以上だった場合、1,000円の送料が無料となります。
まとめ買いをすると1商品あたりのコストパフォーマンスが高くなるためおすすめです。
②プライバシー守る安心梱包
外箱に当サイト名や商品名が記載されることはないため、ご家族や配達員など第三者に内容を知られることは御座いません。

③100%メーカー正規品取り扱い
当サイトの商品は100%メーカー正規品となっており、第三者機関による鑑定も行っております。
商品の破損などがあった場合は再配送などにて対応させて頂きますので、ご連絡頂ければ幸いです。

④いつでも購入可能 処方箋不要
サイト上では24時間いつでもご注文を受けております。
また、お電話によるご注文も受け付けておりますのでネットが苦手な方はお気軽にどうぞ。

⑤商品到着100%
商品発送後はお荷物の追跡状況が分かる追跡番号をご案内させて頂きます。
郵便局には保管期限がありますのでご注意ください。
・自宅配達で不在だった場合の保管期限・・・16日間前後
・郵便局留めとした場合の保管期限・・・7~30日間

⑥コンビニ決済利用可能
ご近所のコンビニにていつでもお支払可能です。
セブンイレブンに限り店舗での機械操作を必要とせず、手続き完了後に表示されるバーコードや払込票番号をレジに提示することでお支払い頂けます。

プロギノバ 2mg x 84錠
9,060円
ポイント:271pt
10,000円以上購入で送料無料
在庫あり

更年期…自分にはいつか来るものと覚悟をしていたつもりでしたが、情緒不安定や異様に汗をかくなどこんなにも辛いものだと思っていませんでした。症状が出るとまたイライラして…というのを繰り返していてかなり参っていたときに義母に進められたのがこのプロギノバでした。1日1回のむだけで症状がかなり抑えられています。薬に頼りたくないという人もいると思うけど、辛いのを我慢するよりかは薬に頼ってもいいってことを教えてあげたいです。
更年期障害は楽になりますが、1箱のお値段が結構しますね。効果はすごく高いので使っていきたいんですが、値段面で不安が残ります。
更年期にみられるホットフラッシュや発汗は多くの人で軽減されます。エストラジオール吉草酸エステルは不足した女性ホルモンを補うことで、体温調整に関わる中枢への影響を和らげ、急なほてりや多汗などの症状を改善する働きがあります。
気分の波が安定してくることがあります。エストロゲンは脳内の神経伝達物質にも関与しており、補充によって軽度の抑うつ気分や情緒不安定が改善するケースも報告されています。ただし、うつ病などの治療とは異なり、あくまで更年期に伴う不調への対応です。
早い人で1〜2週間ほど、一般的には4週間前後で症状の改善を感じるケースが多いです。即効性ではなく、毎日飲み続けることで徐々に変化が現れてきます。
骨密度の低下を抑える効果があります。閉経後にホルモンが減少すると骨が弱くなりますが、この薬はそれを防ぐ目的でも処方されます。骨粗しょう症予防として使われることもあり、医師の判断で長期的に使用されることがあります。
毎日同じ時間に飲めば朝でも夜でも構いません。血中ホルモン濃度を安定させるため、なるべく固定したタイミングで服用することがすすめられています。生活リズムに合った時間帯を習慣化するのが続けやすく、飲み忘れ防止にもなります。
気づいた時点ですぐに飲めば問題ないことが多いです。ただし、次の服用時間が近い場合は1回飛ばしてスケジュールを続けます。1日に2回分まとめて飲むのは避けた方が安全です。
食前・食後のどちらでも大きな差はないとされています。ただし、胃が弱い人は食後に飲んだ方が負担は少ないため継続しやすくなります。また毎回同じタイミングで飲むことで習慣化しやすくなり、飲み忘れの予防にもつながります。
症状が軽くなっても、自己判断で急に中止するのは避けた方がいいです。ホルモン補充療法は段階的に減らしていく必要がある場合が多く、急な中断によって不調がぶり返すこともあります。中止のタイミングや方法については、必ず医師と相談しながら進めるのが基本です。
代表的な副作用として不正出血や頭痛、吐き気などです。軽度で自然におさまることもありますが、症状が強く出たり続いたりする場合は医師に相談が必要です。特に血栓症や乳がんのリスクがある人は使用前に慎重な判断が求められます。
乳がん、子宮内膜がんなどのエストロゲン依存性腫瘍の既往がある人には使用できません。エストラジオールはホルモンに反応する組織を刺激する可能性があるため、がんの再発や増悪のリスクがあると判断されます。既往歴は必ず医師に申告してください。
抗てんかん薬や一部の抗生物質、セントジョーンズワートなどは肝臓の酵素を誘導し、ホルモン剤の代謝を早めて効果を弱めてしまうことがあります。現在服用中の薬がある場合は必ず医師か薬剤師に相談してください。
主な副作用には乳房のはり、不正出血、軽い吐き気、頭痛などがあります。これらはホルモンの変化によって一時的に起こることが多く、通常は継続するうちに落ち着いてくる傾向がありますが、症状が強い場合は医師に相談してください。
| 1日の服用回数 | 1回 |
|---|---|
| 1回の服用量 | 2mg |
| 服用のタイミング | 指定なし |
| 服用間隔 | 毎日同じ時間帯 |
| 商品名 | プレモン | エチニラ | オエストロジェル | プレマリン | メンダポーズ | リノラル | ダーメストリル100 |
|---|---|---|---|---|---|---|---|
| 商品画像 |  |  |  |  |  |  |  |
| 特徴1 | 日本で処方されるプレマリンと同一成分 | 女性ホルモン補充で心身の不調を解消! | 塗布することで更年期の不調を解消 | 更年期のイライラを飲んで治す薬 | ・体に負担をかけずに更年期障害を改善 | ・更年期障害の症状改善に効果的 | ・肌に貼って使用する外用薬タイプ |
| 特徴2 | 価格は先発薬(プレマリン)の約1/6 | 美肌効果も期待できる | 1本で約1ヶ月間使える | 加齢で気になる骨粗しょう症も予防 | ・医薬品よりも長期的な服用に適している | ・副作用も少なく幅広い層の方に人気 | ・信頼性が高いホルモン薬の成分を配合 |
| 内容量 | 0.625mgx112錠 | 0.05mgx112錠 | 80gx1本 | 0.625mg28錠x1箱 | 60錠 | 0.01mgx100錠 | 8mg24パッチx1箱 |
| 価格 | 4,060円 | 5,560円 | 5,760円 | 6,800円 | 4,560円 | 4,250円 | 11,460円 |
| 頻度不明 | |
| 良性、悪性、および詳細不明の腫瘍 | 乳がん、子宮内膜がん |
| 免疫系の障害 | 過敏症反応、遺伝性血管性浮腫の悪化、代謝と栄養障害、ポルフィリン症の悪化、体重の増加または減少、食欲増加、炭水化物耐性の低下 |
| 精神障害 | 不安/抑うつ症状、性欲の低下または増加 |
| 神経系障害 | 片頭痛、頭痛、めまい、疲労、舞踏病、脳卒中 |
| 眼疾患 | 視覚障害、コンタクトレンズ不耐性、心臓疾患、動悸、心筋梗塞、血管障害、高血圧、血栓性静脈炎、静脈血栓塞栓症 |
| 呼吸器、胸部および縦隔の疾患 | 鼻血 |
| 肝胆道疾患 | 胆汁うっ滞を含む胆嚢疾患 |
| 皮膚および皮下組織障害 | 発疹、様々な皮膚疾患(掻痒、湿疹、蕁麻疹、ニキビ、多毛症、脱毛、結節性紅斑、多形紅斑、出血性発疹、肝斑 |
| 筋骨格系および結合組織疾患 | 筋肉のけいれん、脚の痛み |
| 腎臓および尿路疾患 | 膀胱炎のような症状 |
| 生殖器系および乳房疾患 | 子宮筋腫の増大、膣カンジダ症、子宮頸部びらん、膣出血パターンの変化および異常な出血または流れ、突発出血、点状出血(治療を継続する間に出血の不規則性は通常治まります)、月経困難症、膣分泌物の変化、月経前症候群様症候群、乳房分泌物、乳房の圧痛、肥大または痛み |
| 全身障害および投与部位の状態 | 浮腫 |
本製品は海外製のため、期限表記が日本と異なる場合がございます。
パッケージ裏面や側面、シートなどに以下のような表記がされています。
| EXP | 使用期限 例:EXP 12/2025→2025年12月まで使用可 |
|---|---|
| MFG または MFD | 製造日 例:MFG 03/2023 |
| BEST BEFORE | 品質が最も安定している目安日 |
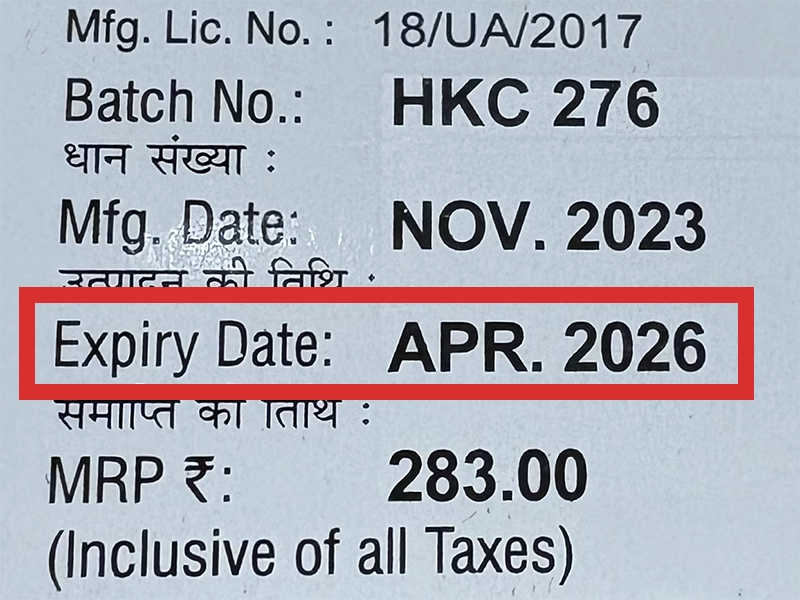
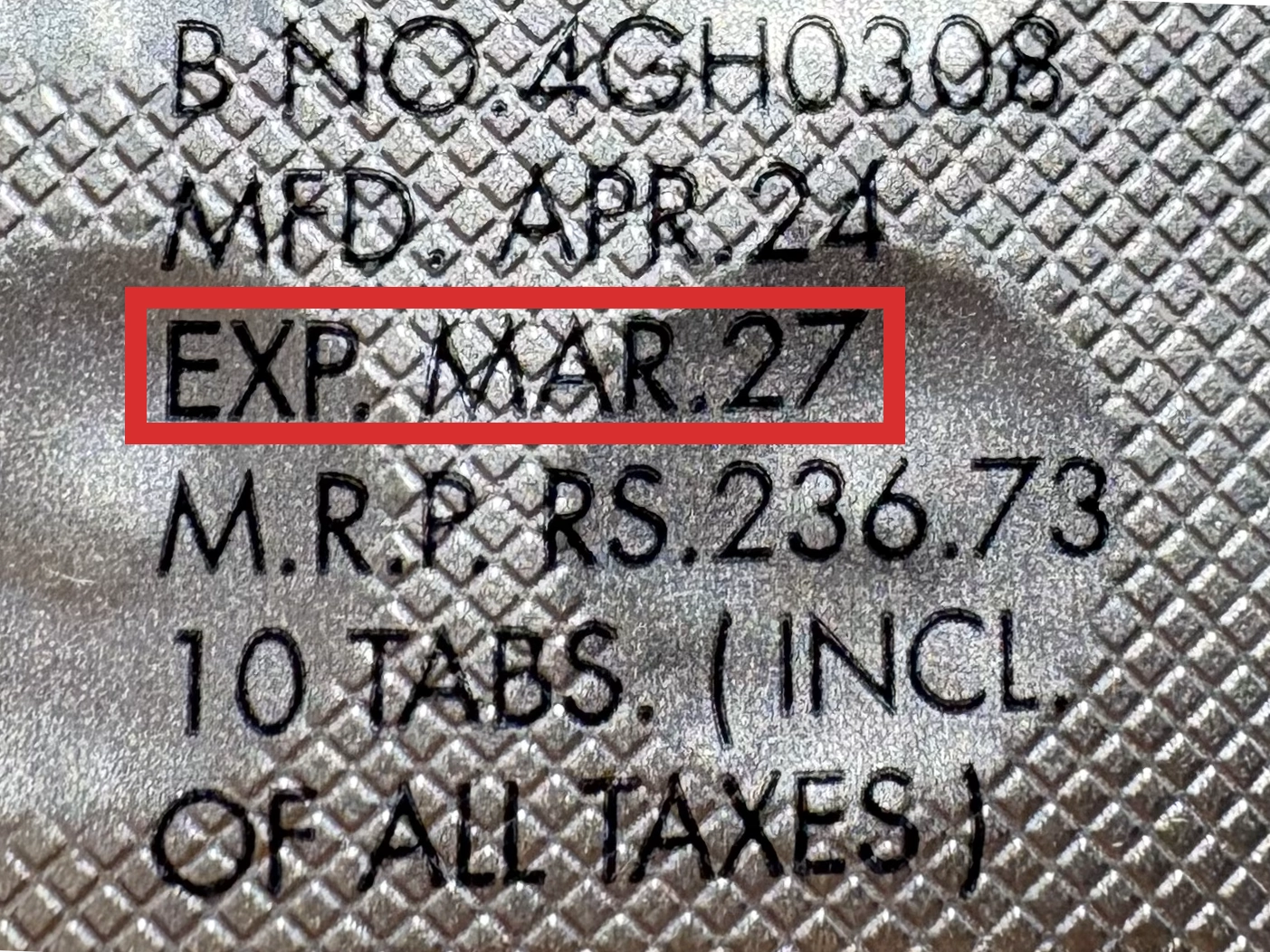
※国や製品により日付の並び(例:月/年、日/月/年)が異なる場合がありますのでご注意ください
EXP(Expiry Date) の表記がなく、MFG または MFDしか記載がないケースがあります。
この場合は MFG(MFD) から2~3年が使用期限の目安です。
※「LOT」や「BATCH」の表記は製造番号であり期限ではありません。

パッケージ例となります。
商品やご注文単位によってはシート単位でのお届けとなる場合が御座います。
外箱に当サイト名や商品名が記載されることはないため、ご家族や配達員など第三者に内容を知られることは御座いません。
50代入った頃から、夜中ひどい寝汗でびしょびしょになったり、頭がぼーっとのぼせたりするようになり、更年期障害を感じることが多くなりました。プロギノバを飲むと、そういう症状は落ち着きます。だけど、吐き気は感じるので、他に体質に合う薬を探しています。
更年期予防としてプロギノバを何度もリピートして服用しています。肌が白くモチモチになってるのが副産物としていいですね。
50代になってイライラすることが多くなり、旦那に理不尽な八つ当たり。優しいので気にしないでくれてるけど、イライラの頻度が多くなって家族からは腫れ物扱いされて、申し訳なくなってきたので、お薬を飲むようになりました。前はプレマリン、今はプロギノバ飲んで、体に合う薬を探しています。
更年期…自分にはいつか来るものと覚悟をしていたつもりでしたが、情緒不安定や異様に汗をかくなどこんなにも辛いものだと思っていませんでした。症状が出るとまたイライラして…というのを繰り返していてかなり参っていたときに義母に進められたのがこのプロギノバでした。1日1回のむだけで症状がかなり抑えられています。薬に頼りたくないという人もいると思うけど、辛いのを我慢するよりかは薬に頼ってもいいってことを教えてあげたいです。
私だけなのかもしれないですが、効果のある症状と効果が薄い症状がある感じです。肩こりや腰痛はかなり軽減されているのですが、不眠やイライラするなど精神的なものにはあまり効果を感じません。
商品口コミの投稿は会員のみ行えるようになっております。
お手数ですが会員ログインの上でご投稿頂きますようお願いいたします。
口コミをご投稿頂いたお客様にはポイントをプレゼントさせて頂いております。
文章のみであれば100ポイント、文章+写真付きのものは300ポイントをプレゼントさせて頂きます。
規約や詳細などはこちらをご確認くださいませ。